Note
Go to the end to download the full example code.
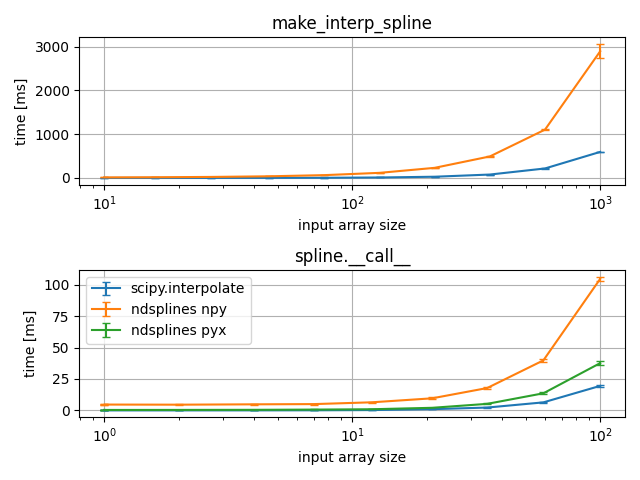
2D ndsplines vs. scipy.interpolate¶
7 import numpy as np
8 import time
9
10 from scipy import interpolate
11 import ndsplines
12
13 import matplotlib.pyplot as plt
14
15 # number of time measurements per input/query size
16 n_iter = 10
17
18
19 def timeit(func, n_iter=1, return_samps=True, **func_kwargs):
20 results = np.empty(n_iter, dtype=np.double)
21 for i in range(n_iter):
22 # gc.collect()
23
24 tstart = time.time()
25 func(**func_kwargs)
26 delta = time.time() - tstart
27
28 results[i] = delta
29
30 if return_samps:
31 return results
32 else:
33 return np.mean(results)
34
35
36 def gen_xyz(sizex, sizey):
37 x = np.pi * np.linspace(-1, 1, sizex)
38 y = np.pi * np.linspace(-1, 1, sizey)
39 xx, yy = np.meshgrid(x, y, indexing="ij")
40
41 zz = np.sin(xx) * np.cos(yy)
42 return x, y, zz
43
44
45 def gen_xxyy(sizex, sizey):
46 x = np.pi * np.linspace(-1, 1, sizex)
47 y = np.pi * np.linspace(-1, 1, sizey)
48 xx, yy = np.meshgrid(x, y, indexing="ij")
49 return xx, yy
50
51
52 # make_interp_spline timing
53 x_sizes = np.logspace(1, 3, 10, dtype=int)
54 t_scipy_build = np.empty((2, x_sizes.size))
55 t_ndspl_build = np.empty((2, x_sizes.size))
56 for i, size in enumerate(x_sizes):
57 x, y, z = gen_xyz(size, size)
58 t_scipy = 10e3 * timeit(
59 interpolate.RectBivariateSpline,
60 x=x.copy(),
61 y=y.copy(),
62 z=z.copy(),
63 n_iter=n_iter,
64 )
65 t_ndspl = 10e3 * timeit(ndsplines.make_interp_spline, x=[x, y], y=z, n_iter=n_iter)
66 t_scipy_build[:, i] = np.mean(t_scipy), np.std(t_scipy)
67 t_ndspl_build[:, i] = np.mean(t_ndspl), np.std(t_ndspl)
68
69 # spline query timing
70 x, y, z = gen_xyz(7, 5)
71 xx_sizes = np.logspace(0, 2, 10, dtype=int)
72 t_scipy_call = np.empty((2, xx_sizes.size))
73 t_ndspl_pyx_call = np.empty((2, xx_sizes.size))
74 for i, size in enumerate(xx_sizes):
75 xx, yy = gen_xxyy(size, size)
76 xxyy = np.stack((xx, yy), axis=-1)
77 spl_scipy = interpolate.RectBivariateSpline(x.copy(), y.copy(), z)
78 spl_ndspl = ndsplines.make_interp_spline((x, y), z)
79 spl_ndspl.allocate_workspace_arrays(size)
80 t_scipy = 10e3 * timeit(
81 spl_scipy, x=xx.copy(), y=yy.copy(), grid=False, n_iter=n_iter
82 )
83 t_ndspl_pyx = 10e3 * timeit(spl_ndspl, x=xxyy, n_iter=n_iter)
84
85 # plot results
86 fig, axes = plt.subplots(nrows=2)
87
88 axes[0].errorbar(
89 x_sizes, t_scipy_build[0], capsize=3, yerr=t_scipy_build[1], label="scipy"
90 )
91 axes[0].errorbar(
92 x_sizes, t_ndspl_build[0], capsize=3, yerr=t_ndspl_build[1], label="ndsplines"
93 )
94 axes[0].set_title("make_interp_spline")
95
96 axes[1].errorbar(
97 xx_sizes,
98 t_scipy_call[0],
99 capsize=3,
100 yerr=t_scipy_call[1],
101 label="scipy.interpolate",
102 )
103 axes[1].errorbar(
104 xx_sizes,
105 t_ndspl_pyx_call[0],
106 capsize=3,
107 yerr=t_ndspl_pyx_call[1],
108 label="ndsplines pyx",
109 )
110 axes[1].set_title("spline.__call__")
111
112 for ax in axes:
113 ax.set_xlabel("input array size")
114 ax.set_ylabel("time [ms]")
115 ax.set_xscale("log")
116 ax.grid()
117
118 axes[-1].legend()
119 fig.tight_layout()
120
121 plt.show()
Total running time of the script: (0 minutes 3.932 seconds)